The last few months I’ve been building out SynthNet’s genetic engine. Each neural structure runs a genetic virtual machine within it that executes genetic instructions obtained from its virtual DNA. Additionally, to make the job of creating this DNA easier, I’ve written a genetic assembly language and a compiler that transforms genetic programs into virtual DNA. The genetic instruction set supports reading and writing any attribute of a structure, including its morphology, ion and protein synthesis, changes in membrane capacitance, growing channels and receptors, changing permeabilities, causing mitosis (soon) or apoptosis, etc. All in all there are around 100 genetic instructions.
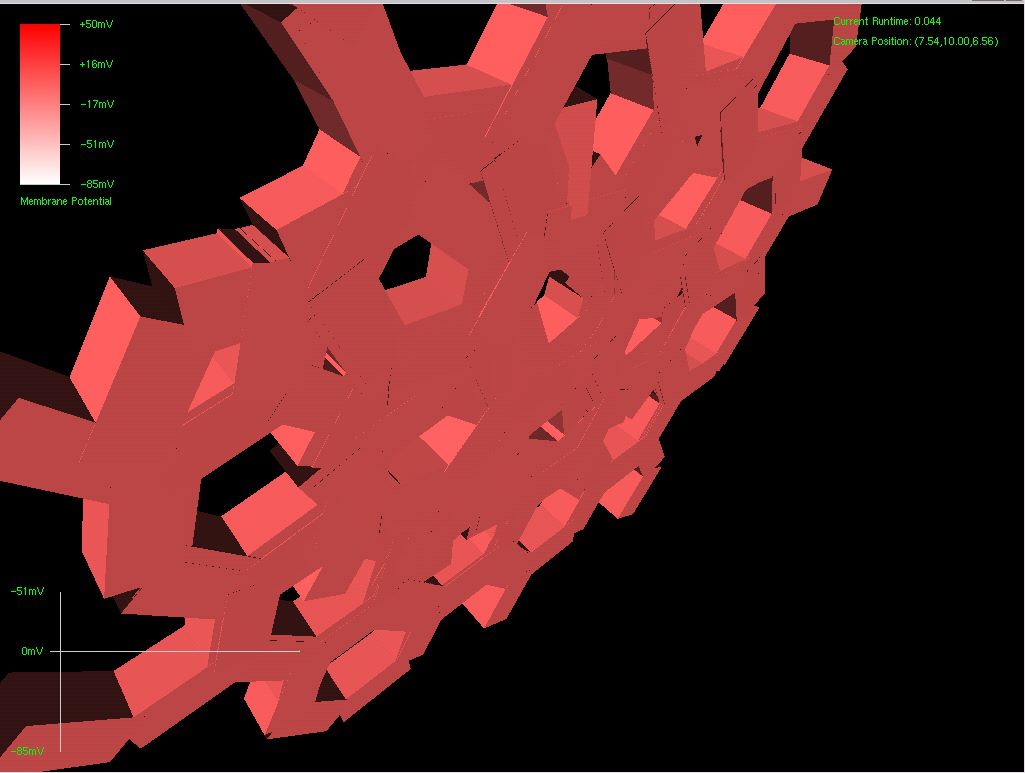
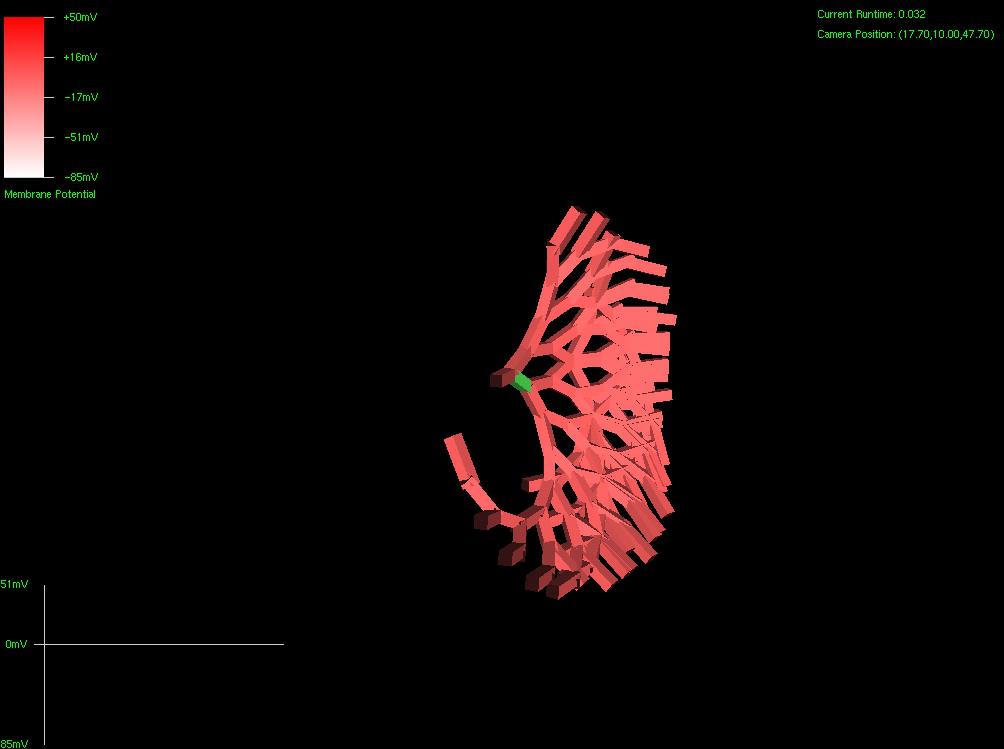
The first picture below is a dendritic arbor I grew using a single strand of virtual DNA . I don’t have enough termination functionality to prevent cancerous growth at this point (soon), but I do have enough to control process growth origination and angle. This was grown with 3 codons worth of DNA – almost nothing. Below that was a snowflake formation, also using 3 codons worth of DNA.

 Hello - and thanks for visiting my site! I maintain ToniWestbrook.com to share information and projects with others with a passion for applying computer science in creative ways. Let's make the world a better and more beautiful place through computing! | More about Toni »
Hello - and thanks for visiting my site! I maintain ToniWestbrook.com to share information and projects with others with a passion for applying computer science in creative ways. Let's make the world a better and more beautiful place through computing! | More about Toni » 





Leave a Reply